phitonex.github.io

Welcome to the Phitonex Github Resource
We are sharing our 40 color flow cytometry data set as a start of this resource. This site intended to show what is possible in leveraging our fluorescent labels, and serve as a reference for opportunities and challenges in high parameter panel design.
40 color Experiment
Pushing the envelope should not mean more difficult data analysis or panel design, and we set out to prove this. Based on a previously shared multicolor panel of 35 colors [1], we started with the 34 commercially available conjugated antibodies and dropped in an 6 additional NovaFluor-conjugated antibodies. Thanks to NovaFluor spectral clarity, performance, and easy conjugation, we were able to immediately plug and play 6 additional colors with limited impact on the current panel, thus pushing our ability to perform deep immunophenotyping without adding complexity in either panel design or data analysis.
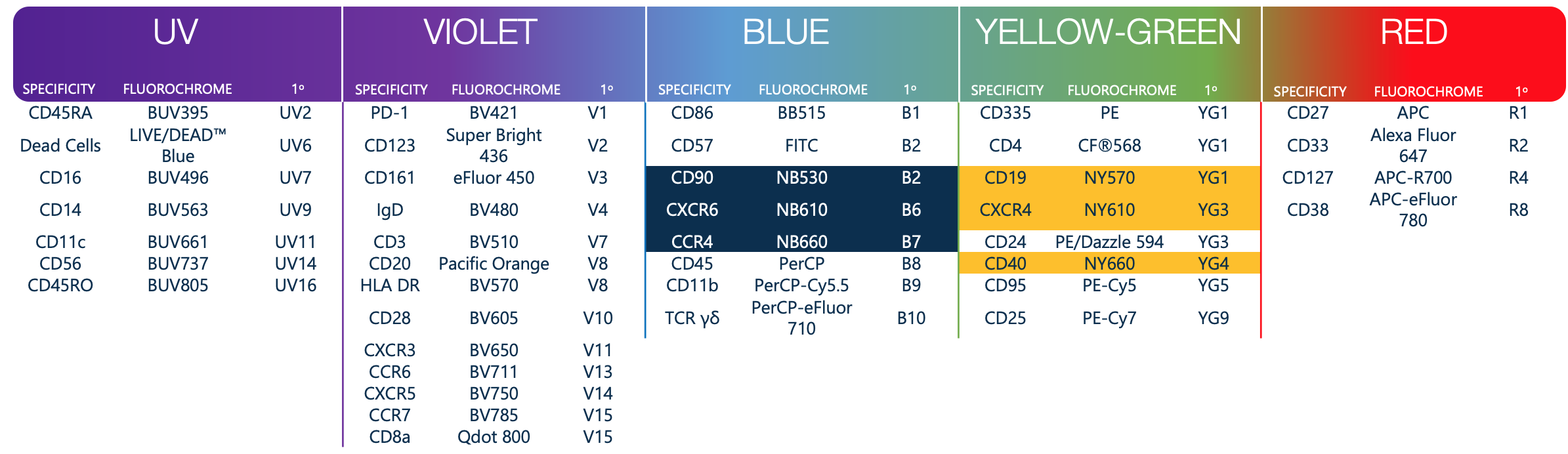
Data Resource
We have shared the data and our analysis under an Attribution-NoDerivatives 4.0 International (CC BY-ND 4.0) creative commons license. [2,3] Data is available here as well as on flowRepository [4].
Purpose
Purpose:Taking a shared 35 color panel [1] we have used the 34 commercially available conjugated antibodies and incorporated our 6 new fluorescent labels. We aimed to assess the “plug and play” ability of 6 NovaFluor labels, as we have designed fluorescent labels that emit at targeted wavelengths with up to 70% less cross excitation than conventional tandem dyes.
Comments
For a deeper discussion of this data set and for all methods, please see our white paper: S.Y. Thomas, C. LaBoda, S. Burrows, D. Daley, A. Stroud, M.D. Stadnisky. “Above and Beyond 40 Colors.” Phitonex, Inc. 2020. or email us info [at] phitonex.com.
QC Measures
Prior to the experiment, single color antibodies were titrated on human PBMCs to determine the optimal staining concentrations for cells. Live-Dead Blue Viability Dye was used to distinguish live cells. NovaBlock™ was used 5 minutes prior to antibody addition to block non-specific cell binding to labels. Single colors were generated on both cells and beads with the most optimal control (cells by default, beads if needed) used for unmixing raw data files. Using the raw data files in SpectroFlo, the full spectra for each label was compared to previously published Cytek spectra to make sure there was no mismatch. The Cytek Aurora instrument was calibrated to default settings prior to acquisition of data.
Conclusion
We have added 6 colors with marginal impact on spread, except for the already difficult BB515-FITC combo. There was near-zero spread added to other channels.
The link below will allow you to download the whole data package which includes:
- Analysis (FlowJo workspace)
- Raw (Unmixed) Data
- Unmixed data
- License
Known issues: When you open the .wsp file in FlowJo, if you do not have the flowSOM plugin installed, you will get an error message. You do not need this to open the workspace, please select:
- “OK” and then
- “Use default” when prompted for the plugin folder location
References
- https://cytekbio.com/pages/aurora#tab-data 2.“Attribution-NoDerivatives 4.0 International (CC BY-ND 4.0).” Creative Commons. https://creativecommons.org/licenses/by-nd/4.0/
- “Attribution-NoDerivatives 4.0 International.” Creative Commons. https://creativecommons.org/licenses/by-nd/4.0/legalcode
- Spidlen J, Breuer K, Rosenberg C, Kotecha N and Brinkman RR. FlowRepository - A Resource of Annotated Flow Cytometry Datasets Associated with Peer-reviewed Publications. Cytometry A. 2012 Sep; 81(9):727-31.
Acknowledgements
We are deeply grateful to the team at Cytek Biosciences who provided us conjugated antibodies, assisted us with SpectroFlo software, and their helpful discussions. We also want to thank the University of North Carolina Cytometry Core especially Nancy Fisher, Ramiro Diz, and Ayrianna Woody for their help and partnership. A big thank you as well to other vendors who assisted us in obtaining reagents, as an experiment like this requires some serious juice. Additionally, a bright and clean thank you is owed to our customer-collaborator (you know who you are, thank you!), Ian Taylor of FlowJo/BD for unmixing support and analysis, Danielle Huppe for her design and critical eye, and Alvin Lebeck for his leadership.
Finally, we are deeply indebted to the numerous customer conversations that informed and pushed our work and our goal to raise the bar in flow cytometry consumables.
Citation
Seddon Y. Thomas, Craig LaBoda, Sean Burrows, David Daley, Aaron Stroud, Michael D. Stadnisky. “Above + Beyond 40 Colors.” Phitonex, Inc. 2020